Discovering
how life works
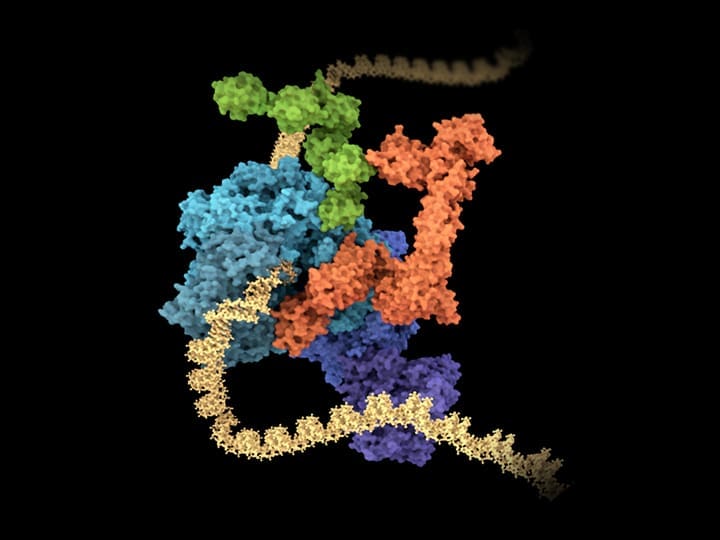
Research that spans the breadth of biology
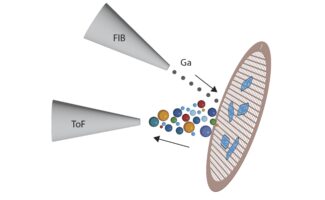
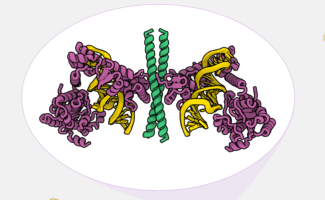
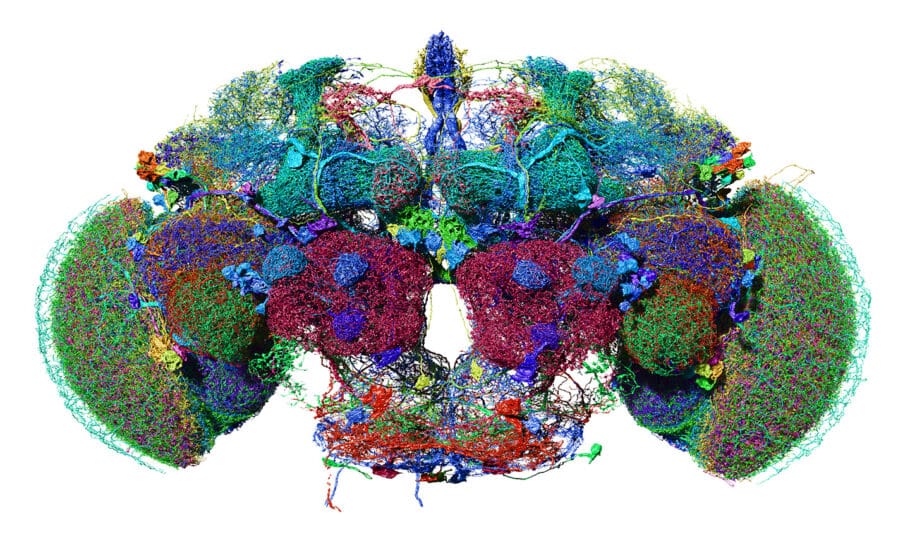
The MRC Laboratory of Molecular Biology (LMB) is a research institute dedicated to the understanding of fundamental biological processes at the levels of atoms, molecules, cells and organisms. The knowledge we provide is crucial to solving key problems in human health.
Latest News

An unsurpassed environment for researchers
The LMB comprises over 50 scientific groups pursuing ambitious research, supported by cutting-edge scientific facilities and generous core-funding by the MRC/UKRI. We occupy an inspiring, purpose-built building on the thriving Cambridge Biomedical Campus.
Upcoming Seminars & Events
Progressing science for over 60 years
Since our origins in 1947, the LMB has pioneered the molecular biology revolution and is a world-leading source of new ideas, scientific advances and technology. Our discoveries have led to highly successful commercialisations and medical advances that have improved human life.